Faster and earlier diagnosis of disease, personalized medicines, and new drug discoveries: there is enormous potential for AI in healthcare to dramatically improve patient outcomes, save lives and help people around the world stay healthier and live longer.
Advances are occurring rapidly and are already having a significant impact. This, in turn, is raising new challenges around the privacy and security of sensitive patient data used to train AI models. Quite rightly, there is growing discussion amongst the public, medical profession and at governmental level about this issue. There is also a clear need for research into improving AI techniques to better protect data privacy.
Now researchers at Stanford University School of Medicine have made a significant breakthrough in the use of Differential Privacy, a key method for securing sensitive data. By using Graphcore IPUs, the Stanford team was able to speed up AI training with Differential Privacy by more than 10x, turning a technique that had been considered too computationally difficult for widespread use, into a viable real-world solution.
Having proven its application using non-privacy sensitive training data, the Stanford team, working with us at Graphcore, now plans to apply its technique to Covid chest CT images, with the aim of uncovering new insights into the virus that continues to impact lives globally.
Data Privacy in Healthcare AI: The Challenge
Using sensitive personal data in AI presents many challenges, but two of the most important are maintaining data sovereignty and preventing the identification of individuals. In each case, sophisticated technological solutions are available – both of which are made more viable by the Stanford research.
Federated Learning
Models trained on large quantities and diversity of patient data, from datasets held by multiple institutions and providers representing different populations around the globe, are more robust, less prone to certain types of biases and ultimately more useful.
Standard machine learning approaches require data to be collated centrally for training. Even where attempts are made to anonymize data, by removing patients’ identifying information, the need to pass this information on to third-party research institutions and organizations has proved hugely problematic. Indeed, there is a growing call for regulation to require that patient data remains in the jurisdiction from which it is obtained.
Federated Learning is part of the answer as it allows AI models to be trained on anonymized patient data without the need to centralize that data. Instead, the model under development is sent out and trained on the data locally.
While Federated Learning is a valuable technique, recent studies have highlighted privacy vulnerabilities around the ability to re-identify anonymous health information by inferring data from a fully trained model, potentially linking it to individuals, or restoring original datasets.
For that reason, use of Federated Learning relies on also advancing the application of Differential Privacy.
Differential Privacy
Differentially private stochastic gradient descent (DPSGD) takes the protection of sensitive data one step further by training a Federated Learning model in such a way that no one can infer training data from it or restore the original datasets.
DPSGD adds noise to anonymized patient data by clipping and distorting gradients for individual training data items. The added noise means an adversary is less likely to uncover the individual patient data used or to restore the original dataset used to train the model.
While offering clear benefits for protecting sensitive data, differentially private stochastic gradient descent (DPSGD) is an under researched area and, until now, has not been applied to large datasets as it is too computationally expensive to use with traditional forms of AI compute such as GPUs or CPUs.
This is the area that the team of radiology researchers specializing in computer vision at Stanford University School of Medicine focused on in their paper, NanoBatch DPSGD: Exploring Differentially Private Learning on ImageNet with Low Batch Sizes on the IPU.
Due to the computational demands, DPSGD is typically applied to and analyzed on small datasets. However, the Stanford team was able to conduct a first analysis on IPU systems using 1.3-million-images taken from the publicly available ImageNet dataset. ImageNet was used in this paper as a proxy for a large image dataset consisting of private data. This work could help overcome current barriers to deploying differential privacy at scale.
Read on for more details of the results with IPUs.
Accelerating NanoBatch DPSGD with Graphcore IPUs
A common approach to accelerate processing is to use micro-batches, where data is processed jointly and the joint gradient gets clipped and distorted instead of individual sample-based gradients. While this accelerates training, analysis shows that this reduces the predictive quality of the resulting model as well as the resulting privacy protection metrics, which essentially defeats its purpose. In fact, running experiments with a micro-batch size of 1, or a “NanoBatch”, shows the highest accuracy.
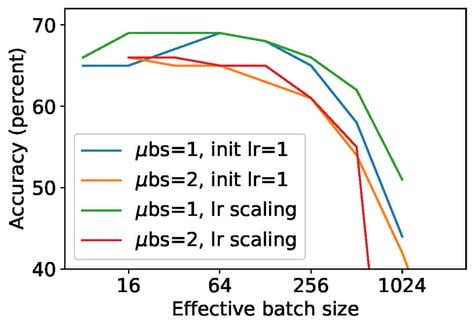
Figure 1: Classification accuracy after 100 epochs of training for various gradient accumulation counts.
We compare micro-batching of size 1 ("NanoBatch") and that of size 2. The effective batch size is the micro-batch size times the gradient accumulation count. The same clipping and noise ratios were used for different effective batch sizes. So, entries with same effective batch size have identical guarantees for privacy protection.
One major reason why NanoBatch DPSGD is not commonly used is that it drastically reduces throughput on GPUs to the extent that running NanoBatch DPSGD ResNet-50 on ImageNet takes days.
By contrast, NanoBatch DPSGD is extremely efficient on IPUs, with results 8–11 times faster than on GPUs, reducing days to hours. For the IPU, the computational overhead from the additional operations required for DPSGD is much lower (10% instead of 50-90%) due to the IPU’s MIMD architecture and fine-grained parallelism enabling far greater processing efficiency.
In addition, privacy preservation and NanoBatch DPSGD requires the use of group norm instead of batch norm, which can be rapidly processed by the IPU but which slows down a GPU significantly. Graphcore Research have recently introduced a new normalization technique, proxy norm, which recuperates batch norm properties for group norm and enhances execution efficiency. This is an interesting future direction to explore.
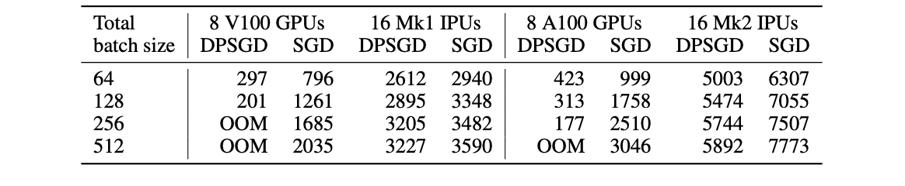
Table 1: Throughput comparison between different hardware for μBS=1.
DPSGD on GPUs uses vmap with the TensorFlow Differential Privacy library. Left part: previous chip generation. Right part: Latest chip generation.
Training NanoBatch DPSGD on ImageNet in 6 Hours
These techniques mean ResNet-50 can be trained on ImageNet in 100 epochs on an IPU-POD16 system in around 6 hours (compared to multiple days on GPUs). An accuracy of 71% was achieved, 5% below the non-private baseline. This was anticipated due to the added noise and while it was better than expected, it remains an area for future research.
For differential privacy, it is common to also report epsilon and delta values. The paper shows an epsilon of 11.4 for a delta of 10-6, which is in a good range. The team has some ideas about how to reduce this further for example, with more aggressive learning rate schedules to decrease the number of epochs.
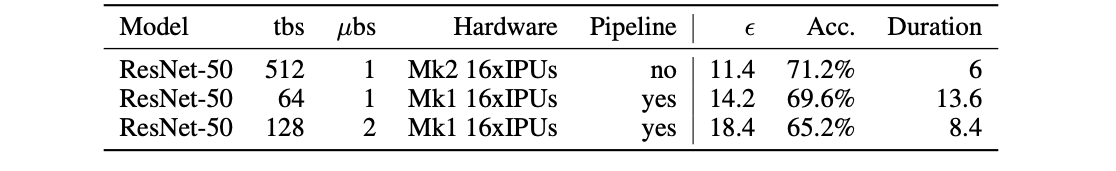
Table 2: Differential privacy results (final ε and accuracy at epoch 100) on ImageNet with different total batch size (tbs) and configurations.
Duration is measured in time it takes to train on 100 epochs in hours.
This research opens up significant opportunities for enhanced privacy for applications in healthcare and many other sectors like financial services where personal sensitive data protection is crucial.
Read the paper
Check out the code